Pirola CJ et. al. (February 2024). Cross talk between the liver microbiome and epigenome in patients with metabolic dysfunction-associated steatotic liver disease EBioMedicine. 101:104996.
The article investigates the interplay between the liver microbiome and epigenome in patients with metabolic dysfunction-associated steatotic liver disease (MASLD). Through analyzing liver microbial DNA composition and assessing histone acetylation and DNA hydroxymethylation levels, the study reveals associations between liver epigenetic factors and the abundance of specific liver bacterial DNAs, suggesting microbial signals may influence the host liver epigenome in MASLD.
Products Used: EpiQuik Nuclear Extraction Kit II (Nucleic Acid-Free), Epigenase HDAC Activity/Inhibition Direct Assay Kit (Colorimetric)
Le D et. al. (February 2024). Neuropathic pain development following nerve injury is mediated by SOX11-ARID1A-SOCS3 transcriptional regulation in the spinal cord Mol Biol Rep. 51(1):281.
The article explores the SOX11-ARID1A-SOCS3 pathway in the modulation of neuropathic pain within the spinal cord following nerve injury. Through experiments in a mouse model of spinal nerve ligation (SNL), researchers observed upregulation of Sox11 post-injury and demonstrated that Sox11 regulates neuropathic pain via transcriptional control of ARID1A, leading to modulation of SOCS3 expression. These findings provide insights into the molecular mechanisms underlying neuropathic pain and suggest potential therapeutic targets for its treatment.
Products Used: EpiQuik Chromatin Accessibility Assay Kit
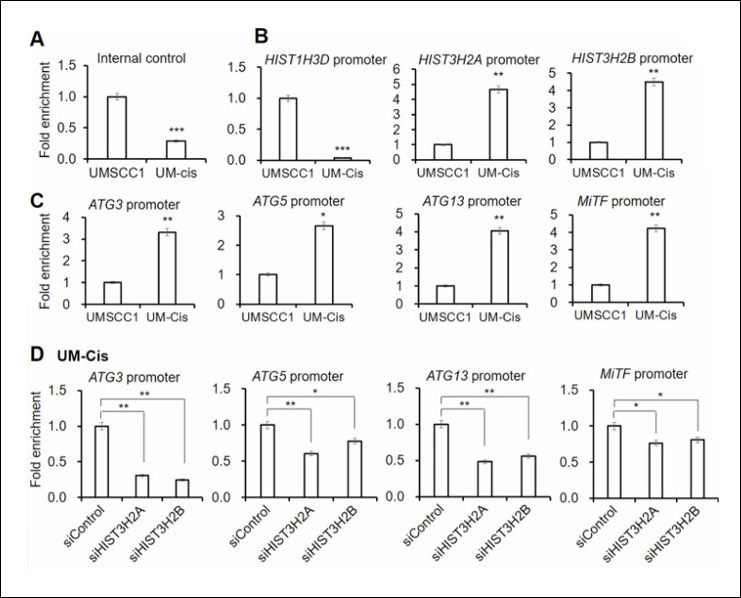
Wu S et. al. (February 2024). The MYC2-PUB22-JAZ4 module plays a crucial role in jasmonate signaling in tomato Mol Plant.
The study investigates the role of the MYC2-PUB22-JAZ4 module in jasmonate (JA) signaling in tomato plants. While COI1 is known to interact with specific JAZ proteins, this study reveals that PUB22 interacts with different JAZ proteins, particularly JAZ4, promoting its degradation via the 26S proteasome pathway. The findings suggest that PUB22-mediated regulation of JAZ4 contributes to various JA responses, highlighting its importance in plant defense mechanisms against environmental stresses.
Products Used: EpiQuik Plant ChIP Kit
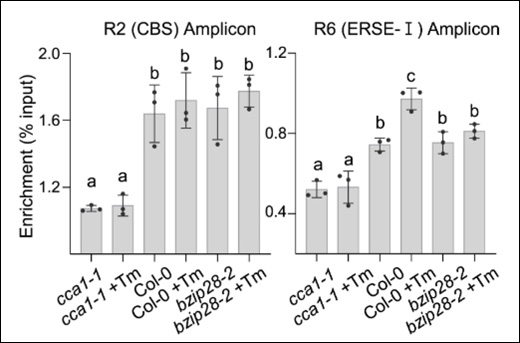
Zhuang H et. al. (February 2024). STAMENLESS1 activates SUPERWOMAN 1 and FLORAL ORGAN NUMBER 1 to control floral organ identities and meristem fate in rice Plant J.
The article investigates the regulatory role of STAMENLESS1 (SL1) in floral organ identities and meristem fate in rice. SL1 interacts with SUPERWOMAN 1 (SPW1) and FLORAL ORGAN NUMBER 1 (DL), influencing stamen development and floral meristem determinacy. Molecular evidence suggests that SL1 acts upstream of SPW1 to regulate stamen development and directly activates SPW1 and FLORAL ORGAN NUMBER 1 (FON1) expression, contributing to floral meristem fate regulation in rice.
Products Used: EpiQuik Plant ChIP Kit
Inoue F et. al. (March 2024). Inhibition of protein arginine methyltransferase 6 activates interferon signaling and induces the apoptosis of endometrial cancer cells via histone modification Int J Oncol. 64(3)
The study examines the role of protein arginine methyltransferase 6 (PRMT6) in endometrial cancer (EC) and its potential as a therapeutic target. Researchers found that PRMT6 inhibition induces apoptosis in EC cells by activating interferon signaling via histone modification, offering a novel approach for EC treatment.
Products Used: Histone H3R2 Dimethyl Asymmetric (H3R2me2a) Polyclonal Antibody
Belle R et. al. (January 2024). Focused Screening Identifies Different Sensitivities of Human TET Oxygenases to the Oncometabolite 2-Hydroxyglutarate J Med Chem.
The study investigates the sensitivity of human TET oxygenases to various inhibitors, including the oncometabolite 2-hydroxyglutarate (2HG). While most inhibitors show similar potencies across TET1-3 and increase cellular 5hmC levels, (R)-2HG exhibits differential inhibition, with TET1 being less affected than TET2 and TET3, suggesting potential implications for tumorigenesis and providing insights for selective inhibitor development.
Products Used: TET1 Protein (Active)
Hou J et. al. (January 2024). Unveiling the mechanism of broad-spectrum blast resistance in rice: The collaborative role of transcription factor OsGRAS30 and histone deacetylase OsHDAC1 Plant Biotechnol J.
The article explores the mechanism behind broad-spectrum blast resistance in rice, highlighting the collaborative action of transcription factor OsGRAS30 and histone deacetylase OsHDAC1. Knockdown of OsHDAC1 boosts blast resistance without altering plant growth, while overexpression of OsHDAC1 heightens susceptibility to Magnaporthe oryzae. This research elucidates how OsGRAS30 inhibits OsHDAC1, elevating histone H3K27ac levels and fortifying blast resistance, offering insights into potential strategies for bolstering crop defense against blast disease.
Products Used: Epigenase HDAC Activity/Inhibition Direct Assay Kit (Colorimetric)
Chen X et. al. (January 2024). Lactylation-driven FTO targets CDK2 to aggravate microvascular anomalies in diabetic retinopathy EMBO Mol Med.
The article investigates the role of FTO, an N6-methyladenosine (m6A) demethylase, in diabetic retinopathy (DR). It explores how elevated FTO expression aggravates microvascular anomalies in DR by promoting angiogenesis, vascular leakage, and retinal inflammation through CDK2 mRNA stability modulation.
Products Used: EpiQuik m6A RNA Methylation Quantification Kit (Colorimetric)
Jung HR et. al. (February 2024). Targeting the m6A RNA methyltransferase METTL3 attenuates the development of kidney fibrosis Exp Mol Med.
The study investigates the role of the m6A RNA methyltransferase METTL3 in kidney fibrosis, a critical mechanism in chronic kidney disease (CKD). By inhibiting METTL3 using siRNA in vitro and the specific inhibitor STM2457 in vivo and in vitro, the study demonstrates a reduction in fibrotic markers and attenuation of kidney fibrosis, suggesting targeting RNA methylation as a potential therapeutic strategy for CKD.
Products Used: EpiQuik m6A RNA Methylation Quantification Kit (Colorimetric)
Caldiran F et. al. (February 2024). Epigenetic insights into Familial Mediterranean Fever: Increased RGS10 expression and histone modifications accompanies inflammation in familial Mediterranean fever disease Gene. 906:148222.
The article investigates the epigenetic regulation of Familial Mediterranean Fever (FMF) and its association with inflammation. RGS10, an anti-inflammatory gene, exhibits decreased expression due to DNA hypermethylation during FMF attacks, while increased levels of specific histone modifications accompany FMF episodes, suggesting significant epigenetic alterations contributing to disease pathogenesis. This research sheds light on the complex interplay between epigenetic mechanisms, immune responses, and FMF, providing potential avenues for therapeutic interventions targeting histone modifications.
Products Used: EpiQuik Circulating Modified Histone H3 Multiplex Assay Kit (Colorimetric)
Jiang L et. al. (February 2024). METTL3-m6A-SIRT1 axis affects autophagic flux contributing to PM2.5-induced inhibition of testosterone production in Leydig cells Sci Total Environ. :170701.
The study explores how long-term exposure to PM2.5 affects testosterone production in Leydig cells through the METTL3-m6A-SIRT1 axis and autophagic flux. In vivo and in vitro models revealed that PM2.5 exposure decreased testosterone levels, disrupted autophagy, and increased m6A modification and METTL3 expression, while knockdown of METTL3 restored autophagy and testosterone production. Activation of SIRT1, downstream of METTL3-m6A, enhanced autophagy and testosterone biosynthesis, suggesting a potential therapeutic target for PM2.5-induced male reproductive injury.
Products Used: EpiQuik m6A RNA Methylation Quantification Kit (Colorimetric)
Fang X et. al. (February 2024). Linc-smad7 is involved in the regulation of lipid synthesis in mouse mammary epithelial cells Int J Biol Macromol. 262(Pt 1):129875.
The article examines the role of Linc-smad7, a long intergenic non-coding RNA, in regulating lipid synthesis in mouse mammary epithelial cells through m6A modification. High expression of Linc-smad7 during pregnancy suggests its involvement in mammary gland development, and experiments demonstrate its influence on lipid formation during mammary epithelial cell differentiation by modulating RNA m6A levels and METTL14 expression.
Products Used: EpiQuik m6A RNA Methylation Quantification Kit (Colorimetric)
Ji X et. al. (February 2024). METTL14 enhances the m6A modification level of lncRNA MSTRG.292666.16 to promote the progression of non-small cell lung cancer Cancer Cell Int. 24(1):61.
The study investigates the role of METTL14 in non-small cell lung cancer (NSCLC) by modulating the m6A modification of the long non-coding RNA MSTRG.292666.16. Elevated levels of METTL14 and m6A modification were observed in NSCLC tissues compared to para-cancerous tissues, and experiments in A549 cells demonstrated that METTL14 knockdown inhibited cell viability, migration, and invasion while promoting apoptosis, further corroborated by in vivo tumorigenicity experiments.
Products Used: EpiQuik m6A RNA Methylation Quantification Kit (Colorimetric)
Qian Q et. al. (February 2024). Acute/chronic triclosan exposure induces downregulation of m6A-RNA methylation modification via mettl3 suppression and elicits developmental and immune toxicity to zebrafish Chemosphere. :141395.
The article explores the effects of Triclosan (TCS) exposure on zebrafish, revealing developmental and immune toxicities, oxidative stress, and reduced m6A RNA methylation levels through downregulation of mettl3 expression. These findings highlight the molecular mechanisms underlying TCS-induced immunotoxicity and ecological risks associated with environmental contaminants.
Products Used: EpiQuik m6A RNA Methylation Quantification Kit (Colorimetric)
Zhao Y et. al. (February 2024). N6 -methyladenosine mRNA methylation positively regulated the response of poplar to salt stress Plant Cell Environ.
The study explores how N6-methyladenosine (m6A) mRNA methylation impacts poplar's response to salt stress, revealing increased m6A levels during this response. It identifies specific transcripts involved in salt response and plant development, with m6A methylation positively regulating gene expression abundance. Additionally, inhibiting m6A writers heightens sensitivity to salt stress, while enhancing the methyltransferase PagFIP37 improves salt tolerance, highlighting m6A methylation's regulatory role in woody plants' response to salt stress.
Products Used: EpiQuik CUT&RUN m6A RNA Enrichment (MeRIP) Kit



 Cart (0)
Cart (0)