m6A ELISA stands as a cornerstone in RNA research, offering a swift, reliable method to quantify N6-methyladenosine in RNA populations. As RNA methylation plays a pivotal role in epigenetic regulation, understanding the intricacies of this technique is of paramount importance.

Understanding the Basics of m6A ELISA
RNA methylation ELISAs, especially m6A ELISA, are a simple and convenient way to rapidly assess global levels of specific methylated RNA modifications, before pursuing more in-depth and costly applications such as qPCR or NGS. Compared to traditional ELISAs, m6A ELISAs are extremely sensitive with much lower detection limits, enabling the assessment of a broader range of sample types.
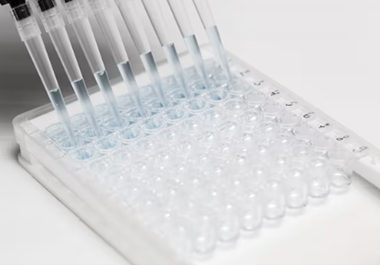 RNA methylation is a reversible chemical modification involved in the epigenetic
regulation of numerous biological processes. It occurs in different RNAs (e.g., tRNA, rRNA, mRNA, tmRNA,
snRNA, snoRNA, miRNA, viral RNA), and distinct catalytic strategies are employed for modifying RNA by a
variety of methyltransferases and demethylases.These methods generally rely on the use of antibodies that
target particular modifications, providing a high-throughput, cost-effective means of analyzing diverse
modified RNA forms.
RNA methylation is a reversible chemical modification involved in the epigenetic
regulation of numerous biological processes. It occurs in different RNAs (e.g., tRNA, rRNA, mRNA, tmRNA,
snRNA, snoRNA, miRNA, viral RNA), and distinct catalytic strategies are employed for modifying RNA by a
variety of methyltransferases and demethylases.These methods generally rely on the use of antibodies that
target particular modifications, providing a high-throughput, cost-effective means of analyzing diverse
modified RNA forms.
While similar to conventional ELISAs in terms of workflow (e.g., sample binding to assay wells, target capture by primary antibody, signal detection with enzyme-conjugated secondary antibody), RNA methylation ELISAs require additional considerations that should be taken into account to ensure reproducibility. Below are some troubleshooting tips for improving the reliability and consistency of your results.
Quality of input RNA
The hydroxyl group at the 2' position of the ribonucleotide sugar moiety renders RNA susceptible to both spontaneous and enzyme-mediated hydrolytic cleavage. Thus, RNA is chemically unstable in comparison to DNA and therefore more demanding to work with. Tissue samples may be stabilized with liquid nitrogen or with a commercial stabilization compound in order to protect RNA prior to extraction. RNA should ideally be extracted from freshly harvested specimens and in a timely manner. If not immediately used, extracts should be stored in single-use aliquots at low temperature (e.g., -80°C).
Methylation from contaminating DNA can also be detected by these assays, as the capture antibody does not discriminate between nucleic acid species. Your RNA sample should be relatively pure, with a 260/280 ratio of 1.9-2.1. The incorporation of DNase treatment into RNA extraction protocols can help minimize DNA contamination. For high-quality total RNA preps, electrophoresis analysis should reveal prominent 18S and 28S rRNA bands, with a 28S/18S band intensity ratio greater than 2. Use fluorometric-based quantification methods such as Qubit, PicoGreen, or SYBR Green, which are more reliable, sensitive, and accurate than photometric approaches like NanoDrop that can overestimate sample concentrations due to the presence of salts, solvents, detergents, proteins, and other contaminants.
Ribonuclease dilemmas
The prevalence of active RNases within samples and in the surrounding environment, especially on hands and surfaces, poses an issue for RNA integrity. In addition to maintaining proper aseptic technique, it should be ensured that RNase-free reagents are strictly used, including water, buffers, pipette tips, glassware, and instruments, and workspaces should be appropriately treated to eliminate RNases.
Choice of primary antibody
Like DNA, RNA contains a diversity of methylation marks (e.g., 5mC, m6A). For immunoassay-based detection, it is hence essential to carefully select a primary antibody that is highly specific for the target methylation and displays no cross-reactivity with off-target modifications. Additionally, the antibody of choice should be validated for use in ELISA application.
Importance of washing
Residual wash buffer can increase background during signal detection. Use manual pipetting during wash steps instead of the common flicking or dumping techniques. Avoid scraping the bottom well surface by guiding the pipette tip down along the sides of the wells, and completely remove the wash buffer from the wells.
Closely monitor color development
RNA methylation ELISAs typically have a greater sensitivity and faster color development time than customary protein ELISAs. To reduce cross-variation between sample replicates, place the replicates vertically within a single column instead of horizontally in the microplate, and add color developer to all wells in each column at the same time using a multichannel pipette.
Know what you’re calculating
Confusion often arises when comparing results obtained with RNA methylation ELISAs and those reported in the literature. For example, some ELISA kits calculate m6A levels as a percentage of methylated A per total RNA (G+A+C+U), and these values will differ from reported percentages for the same sample type that are instead expressed as %m6A per total number of A. Be mindful of this discrepancy, and make literature comparisons accordingly.
Excel with EpigenTek
For quick and accurate quantitation of global RNA methylation via a simple, all-in-one ELISA-like assay, EpigenTek offers the highly cited EpiQuik™ m6A RNA Methylation Quantification Kit. This kit is a complete set of optimized buffers and reagents for the colorimetric quantification of the m6A modification from input samples of purified RNA. Universal positive and negative controls included in the kit are suitable for quantifying m6A in total RNA isolated from any species. A highly specific capture antibody, which displays no cross-reactivity to unmethylated adenosine within the indicated concentration range of the sample RNA, allows for high sensitivity with a detection limit as low as 10 picograms of m6A. Using easy-to-follow steps for convenience and speed, the entire procedure can be completed in just under 4 hours. The strip-well microplate format makes the assay flexible for manual or high-throughput processing, and is available in both 48- and 96-well sizes.



 Cart (0)
Cart (0)